Many bacteria and viruses are protected from the immune system by a thin, hard outer shell — called an S-layer — composed of a single layer of identical protein building blocks.
Understanding how microbes form these crystalline S-layers and the role they play could be important to human health, including our ability to treat bacterial pathogens that cause serious salmonella, C. difficile and anthrax infections. For instance, researchers are working on ways to remove this shell to fight anthrax and other diseases.
Now, a Stanford study has observed for the first time proteins assembling themselves into an S-layer in a bacterium called Caulobacter crescentus, which is present in many fresh water lakes and streams.
Although this bacteria isn’t harmful to humans, it is a well-understood organism that is important to various cellular processes. Scientists know that the S-shell of Caulobacter crescentus is vital for the microbe’s survival and made up of protein building blocks called RsaA.
A recent news release describes how the research team from Stanford and SLAC National Accelerator Laboratory were able to watch this assembly, even though it happens on such a tiny scale:
“To watch it happen, the researchers stripped microbes of their S-layers and supplied them with synthetic RsaA building blocks labeled with chemicals that fluoresce in bright colors when stimulated with a particular wavelength of light.
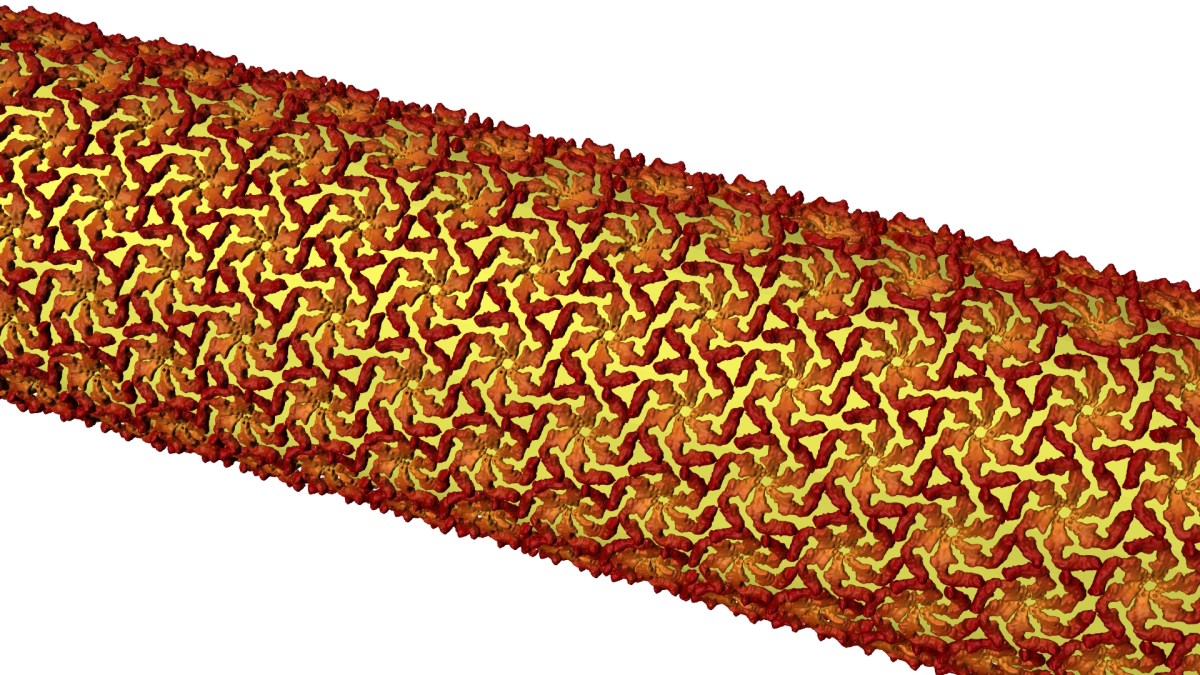
Then they tracked the glowing building blocks with single-molecule microscopy as they formed a shell that covered the microbe in a hexagonal, tile-like pattern (shown in image above) in less than two hours. A technique called stimulated emission depletion (STED) microscopy allowed them to see structural details of the layer as small as 60 to 70 nanometers, or billionths of a meter, across – about one-thousandth the width of a human hair.”
The scientists were surprised by what they saw: the protein molecules spontaneously assembled themselves without the help of enzymes.
“It’s like watching a pile of bricks self-assemble into a two-story house,” said Jonathan Herrmann, a graduate student in structural biology at Stanford involved in the study, in the news release.
The researchers believe the protein building blocks are guided to form in specific regions of the cell surface by small defects and gaps within the S-layer. These naturally-occurring defects are inevitable because the flat crystalline sheet is trying to cover the constantly changing, three-dimensional shape of the bacterium, they said.
Among other applications, they hope their findings will offer potential new targets for drug treatments.
“Now that we know how they assemble, we can modify their properties so they can do specific types of work, like forming new types of hybrid materials or attacking biomedical problems,” said Soichi Wakatsuki, PhD, a professor of structural biology and photon science at SLAC, in the release.
Illustration by Greg Stewart/SLAC National Accelerator Laboratory
This is a reposting of my Scope blog story, courtesy of Stanford School of Medicine.