From table-top experiments to theories about the universe, Berkeley Physics guides diverse efforts in quantum information science
A century ago, science went quantum. In recognition of the progress made since this initial development of quantum mechanics, 2025 is being celebrated as the International Year of Quantum Science and Technology.
According to the quantum view of nature, matter absorbs energy in tiny discrete packets. And seemingly disconnected objects can be entangled, their properties correlated by intangible links—even if they are light years apart.
Initially, quantum mechanics was something specialized physicists thought about, including posing philosophical questions like whether Schrödinger’s quantum cat is simultaneously dead and alive. Beginning in the 1970s, however, these theories were tested by experiments that proved entanglement is real and fundamental to nature. The first definitive proof of quantum spookiness took place experimentally in Birge Hall and was recognized by the 2022 Nobel Prize in Physics. Since the 1990s, lots of quantum research focuses on how to develop quantum technologies.
“Quantum mechanics has transformed from a philosophy to an engineered product—that’s a very big deal,” says Irfan Siddiqi, Berkeley Physics professor and chair. “Bringing these technologies to the market to change our everyday world requires people from industry, academia, and venture capital to come together. Berkeley is at the center of this convening effort, because it offers a unique entrepreneurial ecosystem with broad and diverse efforts in quantum information science.”
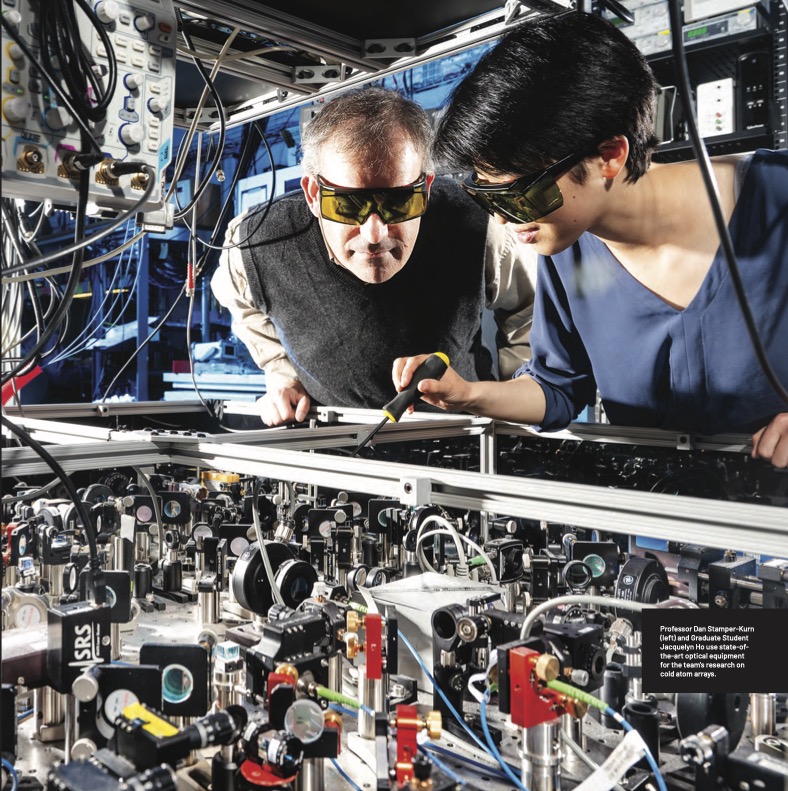
Cutting-edge researchers at Berkeley Physics lead investigations of all competing technologies for creating and manipulating qubits, the basic unit of information in quantum computing—including research on cold atom arrays by Professor Dan Stamper-Kurn, superconducting circuits by Siddiqi, trapped ions by Professor Hartmut Haeffner, and advanced quantum materials by various faculty. Berkeley also has world-leading theoretical physicists such as Professor Raphael Bousso, mathematicians, and computer scientists exploring fundamental aspects of quantum information. We highlight only a few of these efforts here.
Table-Top Experiments with Ultracold Atoms
Although quantum research has made amazing progress over the last 100 years, we’re still really far from building a quantum computer that you could use at home, according to Stamper-Kurn.
“We also still don’t know if a quantum computer is good for much. We know they can factorize large integers and simulate quantum mechanical systems, which classical computers will never be able to do at the same scale,” he explains. “We take it on faith that this is just the tip of an enormous ice berg, but that isn’t proven yet.”
However, needing to tackle these big challenges means Berkeley physicists have tons of interesting research ahead, including work led by Stamper-Kurn at the NSF Challenge Institute for Quantum Computation (CIQC).
“We use the term institute because we’re trying to do something more curiosity-driven and longer-term than a single research project with a specific deliverable. I think academia’s role should be to know where the road is taking us, but to also be willing to look at the side roads—find technology breakthroughs by stumbling around and bring them to the fore,” say Stamper-Kurn.
At CIQC, his multidisciplinary team of physicists, computer scientists, mathematicians, chemists, and engineers pursue three overarching goals. The first is to engineer large-scale coherent quantum systems, which is where Stamper-Kurn and Siddiqi overlap a lot. The second goal is to realize the quantum computer and figure out what it can do, which leads CIQC researchers to interact with a wide range of physicists and computer science theorists. The third goal is to use concepts of quantum information science to understand the natural world, and that’s where Stamper-Kurn learns from Bousso.
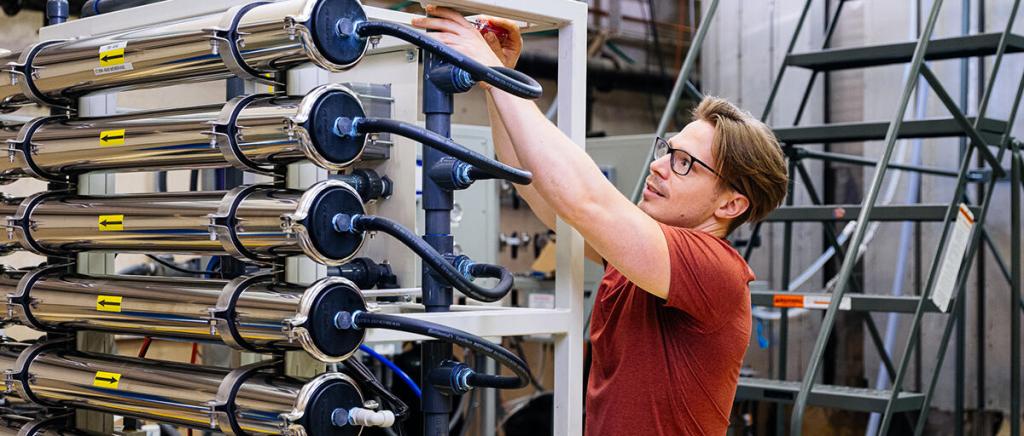
More specifically, Stamper-Kurn’s group performs table-top experiments using light and ultracold atomic gases—perhaps the coldest matter in the universe. At temperatures below one nano-kelvin, noise is “ironed out” and the quantum mechanical properties of these atoms are uniquely accessible and visible. Their goal is to image, with high resolution, individual cold atoms to provide extremely sensitive quantum measurements.
In the E6 ultracold atoms lab, his team studies how a single or array of ultracold atoms interact within the optical field of a high-finesse optical cavity, where light shined into the cavity bounces back and forth thousands of times between two end mirrors. Since light is trapped in the cavity for so long, it interacts very strongly with any atoms inside.
Optical tweezers, consisting of highly focused laser beams sent through a high-resolution lens, are used to control the atoms. They precisely trap atoms in different locations of the cavity or manipulate the state of an individual atom. The team also uses the high-resolution lens to collect light from the atoms as it comes out of the cavity, which then passes through an imaging system and is focused onto a camera.
“We use arrays of optical tweezer traps to position arrays of neutral atoms within a cavity, achieving unprecedented control of the position, internal state, and optical response of each individual atom,” says Stamper-Kurn.
“This setup has allowed us to realize breakthroughs in quantum computing, including our recent rapid mid-circuit measurement of atom tweezer arrays,” he explains. “We use the very strong interaction between atoms and light to read out the state of an atom very rapidly—about 1000 times faster than previous experiments using optical tweezers. But real applications require measuring part of a quantum processor while the rest of it keeps computing. Our technique is also very selective, where only the atom we measured was disturbed.”
Additionally, CIQC supports the entire campus in its development of quantum science, including funding and organizing “Quantum Gatherings” attended by students, postdocs, and faculty from various departments. Held every other Friday, someone from the broader Berkeley quantum community gives a provocative 20-minute spark talk followed by an hour-long discussion.
A Berkeley undergraduate student who attended the gatherings sent Stamper-Kurn a note saying, “Thank you for sparking my interest in quantum science. I came for the pizza, and I stayed for the science.”
These gatherings are the “sweet place” where Stamper-Kurn, Siddiqi, Bousso, and their groups interact. “We can be seen chewing pizza and talking about physics every two weeks outside of Campbell Hall. We understand this is a long-term and necessarily multidisciplinary effort to create an ecosystem for quantum information science,” Stamper-Kurn says.
Full-stack Applications with Superconducting Systems
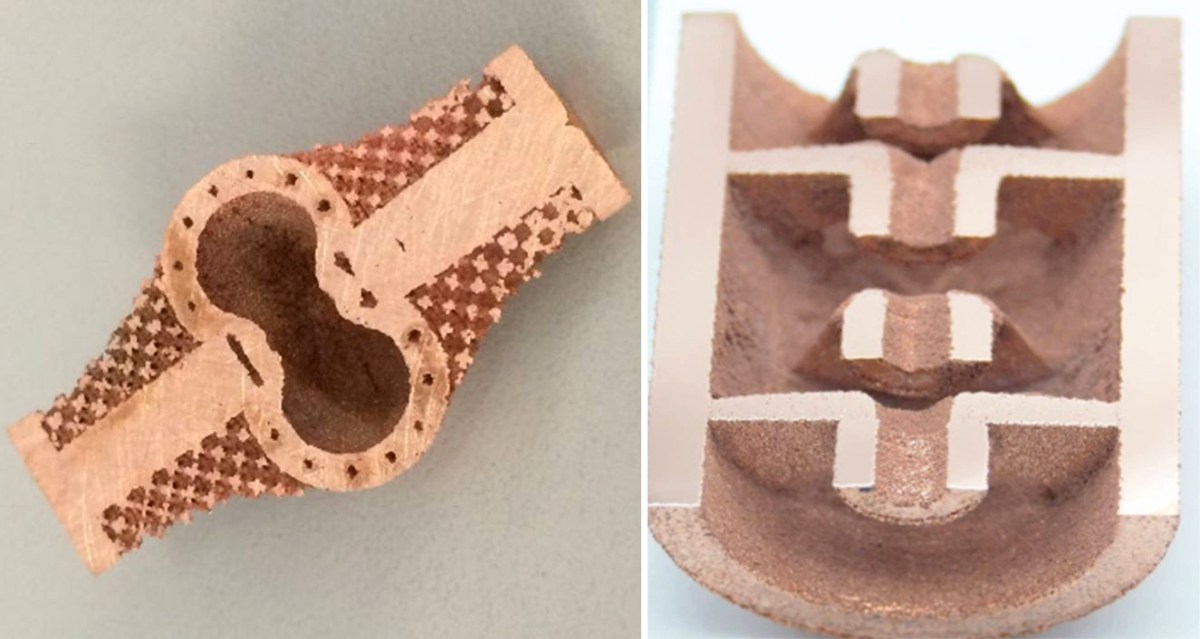
While Stamper-Kurn performs table-top cold atom experiments, Siddiqi’s Quantum Nanoelectronics Lab (QNL) works on a larger scale with full-stack quantum computers using superconducting devices. This comprehensive program includes developing materials and processors and integrating them into platforms with controls and applications layers.
QNL has its own clean room and cryogenics for building and operating large superconducting systems on campus. Siddiqi is also the director of the Advanced Quantum Testbed at Berkeley Lab, which offers white-box access to superconducting quantum computers through a user program.
“My large team has fingers in many pots. We have many quantum computers running based on a range of superconducting hardware,” says Siddiqi. “We’re still at a stage where we haven’t figured out what the right qubit is, how to use it, and how to connect it—before scaling up.”
Siddiqi focuses on superconducting platforms because they are open quantum systems that naturally exchange information with the environment. These systems already have many “knobs,” so the challenge is figuring out how to interface with them properly, he says.
“This is a new chapter in quantum mechanics, originally a theory of closed quantum systems. We’re now looking at how to measure and engineer open quantum systems, both for computation but also to explore the most fundamental questions,” says Siddiqi.
One technology QNL is developing is a quantum information processor that uses entangled three-level quantum systems, or qutrits.
In a classical computer, a bit is a 0 or 1, represented by on-off switches in hardware. But what if the switch could exist as a coherent superposition of 0, 1, and the states in between simultaneously—think, the Schrödinger cat is both alive, dead, and a zombie? And what if the switches in a computer could consult each other before outputting a calculation?
For example, a quantum computer with an entangled array of N two-level qubits can represent 2N possible states—far more than the N states of a classical computer—enabling it to perform calculations on many possibilities in parallel. And that’s just a binary system.
A three-level qutrit system offers an even larger and more connected computational space that increases with 3N. But as you add levels, it becomes more challenging to control and entangle them while making them robust against undesired noise, crosstalk, and errors.
Siddiqi’s team tackled these challenges, experimentally implementing faster, flexible, and tunable microwave-activated entanglement with three or more levels.
Using their new approach for entanglement, Siddiqi’s team developed two types of high-quality two-qutrit gates, a controlled-Z gate and a controlled-Z inverse gate—decreasing the gate’s error rate by a factor of four over previous efforts, leading to higher computational performance.
“In a static case, when the qutrits are not driven by electromagnetic fields, they’re not coupled. But if we periodically drive the qutrits, like a pendulum back and forth, we can create entanglement on demand—and that’s a powerful tool,” says Siddiqi. “That obeys different rules, so we’re able to create this N-body entanglement.”
This significant step forward for operating multi-qutrit devices will help pave the way for a deeper understanding of ternary quantum logic, which can encode more information in quantum processors than qubits.
More generally, Siddiqi is also leading efforts to create a horizontal ecosystem for quantum technologies at Berkeley and beyond, including founding the consortium Berkeley Quantum Works, which fosters partnerships between the device fabrication and measurement facilities housed on the Berkeley campus and startup-stage companies who make quantum processors, components, and software.
Theories on Quantum Information in the Universe
Berkeley’s quantum efforts go beyond developing and using advanced technologies for quantum computation. Colleagues like Bousso are investigating quantum on the scale of the universe.
“Raphael is thinking about the universality of quantum information as a currency to describe the physical world—both in the celestial, gravitational, and cosmology sense and in the computing sense,” describes Siddiqi.
The Bousso group uses diverse tools and techniques to investigate the increasingly important intersection of quantum information and quantum gravity.
The field of quantum gravity seeks a unified theory of our universe that includes both quantum mechanics and general relativity. It generally focuses on the interplay between spacetime geometry, quantum information theory, and relativistic field theory. However, according to Bousso, this theoretical challenge is really about how quantum information and gravity are intertwined.
“It’s about hunting the thrill of quantum gravity. Once we really understand it, there’s every reason to believe it’s going to revolutionize how we think about the world,” says Bousso, who is the Chancellor’s Chair in Physics.
Much of Bousso’s research focuses on black holes, which are massive deformations of the shape of space and time.
“Black holes are where we can sharpen what we don’t understand about quantum gravity—where we can turn vague confusions into a sharp paradox,” says Bousso. “For example, in the black hole information paradox, either black holes destroy quantum information in which case quantum mechanics is terribly wrong or you can’t cross a black hole’s horizon in which case general relatively is terribly wrong. These paradoxes force us to make stark choices, like which principle to give up, and that blossoms progress.”
Another fundamental question concerns a black hole’s singularity, the point of infinite density and gravity within its event horizon where all concepts of time and space break down. Can you evade that singularity with quantum corrections?
Some theoretical models of quantum gravity suggest that the singularity inside a black hole is not a true point of infinite density, but instead a region where spacetime undergoes a bounce, potentially transitioning to a white hole. This could resolve the information paradox. Other theorists suggest that the universe is infinitely cyclically bouncing between contraction and expansion, replacing the “big bang” with a “big bounce” theory.
Bousso recently ruled out these sorts of cyclic cosmologies with his Robust Singularity Theorem, showing that you cannot get through such bounces.
“People have tried playing with many ideas like this, formulated at various levels of rigor and with varying levels of plausibility. What’s nice is that my theorem rules them out regardless of the details,” he says.
His Robust Singularity Theorem expanded the validity of the Penrose-Wall singularity theorem, which showed singularities really exist and they arise in many situations in general relativity when spacetime contains a trapped surface and specific conditions are met. Whereas the Penrose-Wall theorem applied to a mathematically idealized case, Bousso’s more general theorem represents a big advance.
Bousso’s work largely focuses on black holes and gravity, but he’s ultimately exploring fundamental questions of entanglement and quantum information. And he’s using ideas from other fields, including quantum information theory and quantum communication protocols, that were motivated by seemingly entirely different problems.
And what he learns about these fundamental questions has the attention of his experimental colleagues. In addition to working with Berkeley Physics experimentalists, he is part of the GeoFlow consortium that includes experimentalists at Stanford and Duke universities.
“The depth at which really sophisticated concepts in quantum information theory are baked into gravity has led us to have something to say to people who are interested in benchmarking their quantum computing platforms,” explains Bousso. “It’s a whole new ecosystem we’re developing, connecting people like me studying quantum gravity to condensed matter theorists and even atomic, molecular, and optical physics experimentalists. It’s fun and has been the best part of my career.”
Irfan concludes, “At first the people from different fields working on quantum spoke different languages, but now they are all very fluent.”
This is a reposting of my magazine feature, courtesy of UC Berkeley’s 2025 Berkeley Physics Magazine.